BioExcel: Electronic Interaction Phenomena: Proton Dynamics and Fluorescent Proteins
Short description
Proton Dynamics
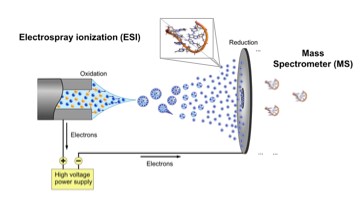
Mass spectrometry has revolutionized proteomics, i.e. the investigation of the myriad of protein/protein interactions in the cell. By using a very small sample content of proteins (usually in the micromolar concentration), a powerful implementation of the technique, so-called ionization/ion mobility mass spectrometry (ESI/IM-MS), can measure stoichiometry, shape and subunit architecture of protein and protein complexes in the gas phase.
Fluorescent proteins are the backbone of many high-resolution microscopy techniques. To improve resolution, these proteins may require optimization for the specific conditions under which the imaging is carried out. While directed evolution works well if a single property needs to be optimized, optimizing multiple properties simultaneously remains challenging. Furthermore, this approach provides only limited physical insights into the process that is the target of the optimization. In contrast, computational chemistry provides a route to obtain such insights from first principles, but because such calculations typically require a high level of expertise, automatic workflows for in silico screening of fluorescent proteins are not yet generally available. To overcome this limitation we developed a user-friendly workflow for automatically computing the most relevant properties of fluorescent protein mutants based on established atomistic molecular dynamics models.
Fluorescent proteins are the backbone of many high-resolution microscopy techniques. To improve resolution, these proteins may require optimization for the specific conditions under which the imaging is carried out. While directed evolution works well if a single property needs to be optimized, optimizing multiple properties simultaneously remains challenging. Furthermore, this approach provides only limited physical insights into the process that is the target of the optimization. In contrast, computational chemistry provides a route to obtain such insights from first principles, but because such calculations typically require a high level of expertise, automatic workflows for in silico screening of fluorescent proteins are not yet generally available. To overcome this limitation we developed a user-friendly workflow for automatically computing the most relevant properties of fluorescent protein mutants based on established atomistic molecular dynamics models.
Results & Achievements
Proton Dynamics
The project is in progress. It has started in December 2020. So far, we have investigated the nucleotide in water. We found a stable hairpin structure consisting of a short B-DNA segment and a d(GpApAp) triloop. Labile contact ion pairs were observed between the heptanucleotide backbone and ammonium ions, which are important for proton dynamics upon desolvation. These calculations are used as a starting point for QM/MM simulations and will be afterwards complemented by investigation of the biomolecule in the gas phase.
Fluorescent Proteins
To illustrate what can be done with the workflow, we have computed the absorption and fluorescence spectra, as well as the folding and dimerization free energies for five variants of Aequorea victoria Green Fluorescent Protein. Other important properties, such as excited-state reactivity, can be computed as well, but require significantly more computational resources. To demonstrate also this aspect of our workflow, we computed an excited state trajectory of the enhanced GFP (EGFP) variant. Because computational power, as well as the accuracy of simulation models, are expected to improve further, we anticipate that our workflow can eventually provide the fluorescence microscopy community with a powerful and predictive tool for tailoring fluorescent proteins to the specific conditions of an experiment.
Objectives
Proton Dynamics
We will use massively parallel hybrid QM/MM approaches to investigate proton dynamics during the desolvation process of biomolecules during mass-spec experiments. The approaches include the previously developed interface (coupling GROMACS to CPMD) to the new interface (coupling GROMACS to CP2K) developed by BioExcel. We will focus on the heptanucleotide d(GpCpGpApApGpC). Comparison with the available experimental data will allow us to establish the accuracy of the methods used. This project will illustrate computational scaling, performance, and efficiency of BioExcel’s QM/MM codes, run on a highly relevant biological problem.
Fluorescent Proteins
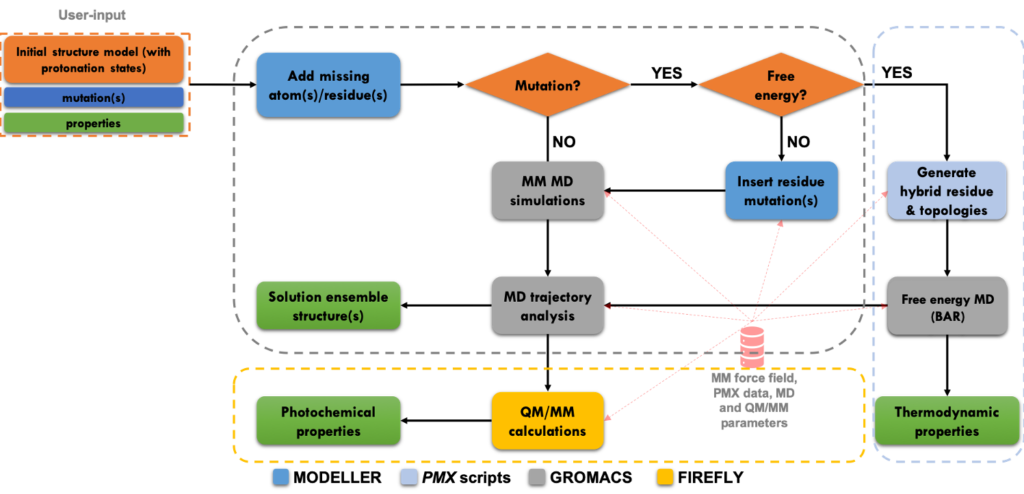
We developed a user-friendly workflow for automatically computing the most relevant properties of fluorescent protein mutants based on established atomistic molecular dynamics models. In particular, the workflow combines classical molecular dynamics, with free energy computations and hybrid quantum mechanics / molecular mechanics (QM/MM) simulations to compute the effects of mutations on the key properties of these proteins, including absorption and emission wavelengths, folding free energy, oligomerization affinities, fluorescence and switching quantum yields.